Research Project
CytoWave
An orthology-aligned foundation model for single-cell genetic perturbation responses across studies and species.
Overview
Single-cell genetic perturbation experiments provide direct measurements of how cells respond to gene perturbations, but these datasets remain fragmented across studies, species, and experimental protocols. As a result, most existing single-cell foundation models are pretrained primarily on unperturbed data and treat perturbation modeling as a downstream task.
CytoWave addresses this limitation by unifying large-scale cross-species perturbation datasets through orthology alignment and pretraining directly on perturbation data. The model is designed to learn transferable representations of perturbation-induced cellular state changes, enabling more faithful modeling of transcriptomic responses under genetic intervention.
Key Idea
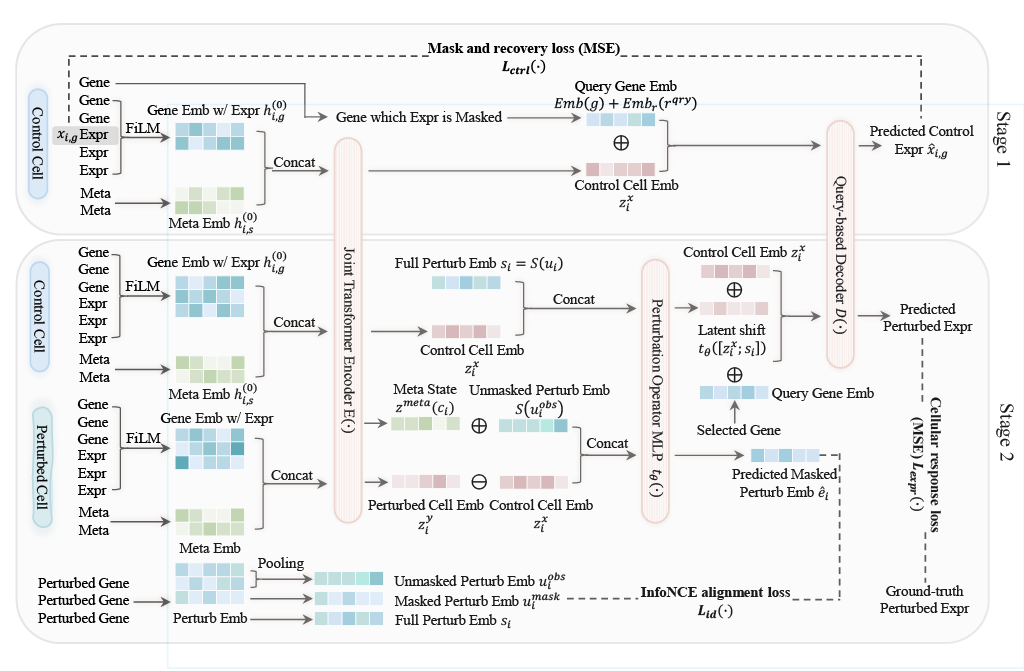
CytoWave is trained in two self-supervised stages. In the first stage, the model learns control cell representations through masked gene expression recovery. In the second stage, it learns a deterministic perturbation operator by predicting post-perturbation cellular responses and aligning perturbation effects contrastively in latent space.
This design allows the model to capture perturbation effects as structured transformations of cellular state, rather than treating perturbation prediction as a purely supervised downstream mapping. Across benchmarks, CytoWave supports accurate perturbation response prediction and reliable inference of pathway-level activity changes after perturbation.