Research Project
GoBERT
Learning gene function relationships from Gene Ontology graphs using BERT-based representation learning.
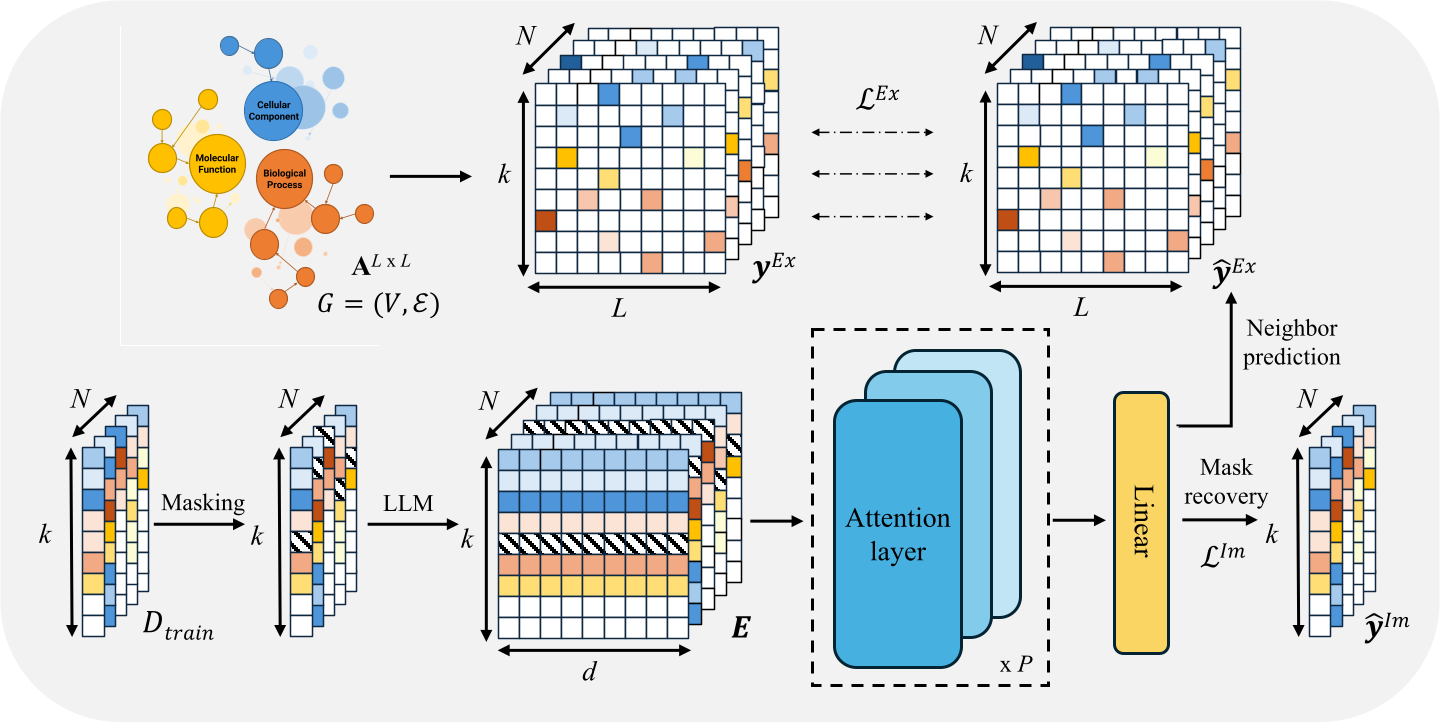
Overview of GoBERT for modeling functional relationships in the Gene Ontology graph.
Overview
Understanding gene functions is essential for biological discovery, drug development, and disease research. However, identifying new gene functions typically requires extensive wet-lab experimentation, which is costly and time-consuming. Computational approaches aim to accelerate this process by predicting gene functions automatically.
Most existing methods rely on biological features such as protein sequences, structural information, or protein family annotations. In contrast, GoBERT focuses on the relationships between functions themselves by modeling the structure of the Gene Ontology (GO) graph and existing functional annotations.
Key Idea
GoBERT formulates gene function prediction as a representation learning problem over the Gene Ontology graph. The model is trained using two complementary self-supervised objectives. A neighborhood prediction task captures explicit functional relationships between GO terms, while a specified masking and recovery task enables the model to learn implicit functional patterns within the ontology.
After pretraining, GoBERT can infer novel gene functions based on existing functional annotations. Experiments, ablation studies, and biological case analyses demonstrate that GoBERT effectively captures functional dependencies and supports accurate prediction of previously unannotated gene functions.