Research Project
UniEntrezDB
A unified database integrating Gene Ontology annotations across biological resources through the Entrez gene system.
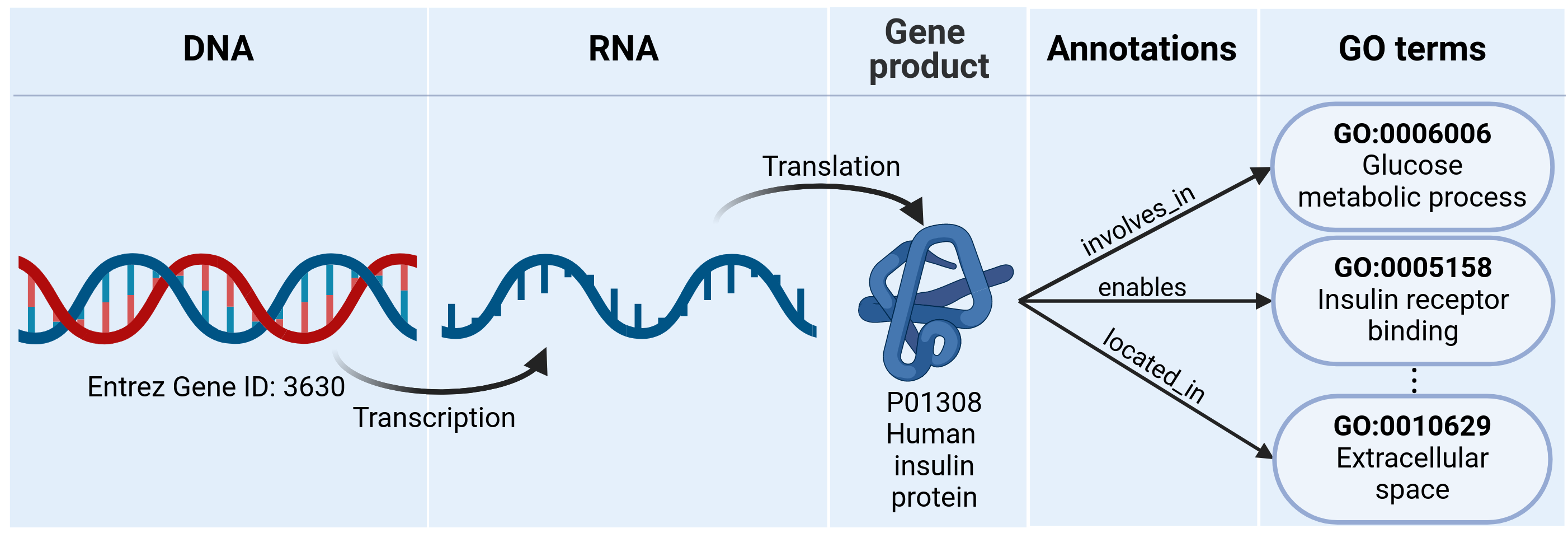
Overview of UniEntrezDB integrating Gene Ontology annotations across multiple biological databases.
Overview
Gene function prediction plays a central role in protein characterization, drug discovery, and cancer genomics. However, biological knowledge relevant to gene functions is distributed across many databases, often using different identifiers for biological entities. This fragmentation makes it difficult to integrate datasets and build reliable machine learning models.
Gene Ontology (GO) provides a structured vocabulary organized as a directed acyclic graph to describe gene functions. Despite its widespread use, GO annotations are scattered across multiple biological resources without a unified identifier system, creating challenges for large-scale data integration.
Key Idea
UniEntrezDB addresses this problem by unifying Gene Ontology annotations from representative public biological databases through the Entrez gene system. The database consolidates heterogeneous biological resources into a single standardized framework, enabling consistent mapping between genes, gene products, and functional annotations.
In addition to the integrated database, UniEntrezDB introduces four benchmark tasks designed to evaluate representation learning from biological object information. Experiments demonstrate that incorporating unified gene functional knowledge can significantly benefit a wide range of downstream biological modeling tasks.